General Research Interests
To understand the organizing logic of living systems.
Check out my YouTube channel for research related videos.
Current Research
Molecular mechanisms and functional impacts of cytoplasmic self-organization
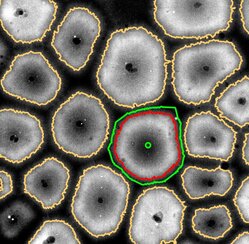
How do cytoplasmic components assemble into biologically functional spatial architecture? To answer this question, we need to observe de novo assembly of spatially organized cytoplasm. This is not possible to do in living cells as they inherit their spatial organization from mother cells instead of building it from scratch. We have discovered that completely scrambled frog (Xenopus laevis) egg extracts, which contain all essential components of the cytoplasm at their native concentration, are able to carry out de novo assembly of spatially organized, biologically functional cytoplasm in vitro (Read more in Cheng and Ferrell, 2019). This unique property makes egg extracts an ideal model system for understanding the mechanisms of de novo cytoplasmic assembly.
Using the Xenopus egg extract system, we are exploring the functional impacts of cytoplasmic organization. We discovered that diffusion of protein-sized molecules (but not molecules much smaller or larger in size) becomes much faster in spatially organized cytoplasm compared to scrambled cytoplasm. Thus, self-organization could in principle enable a broad range of diffusion-limited biochemical reactions that would otherwise be too slow to be physiological (Read more in Huang, Cheng and Ferrell, 2022). We seek to further uncover the fundamental design principles of living cellular systems by dissecting the mechanisms of self-organization in frog egg cytoplasm. These principles will be instructive to the engineering of synthetic living systems, and may shed light on the origin of cellular life. My lab is currently investigating:
(1) How is the microtubule (MT) and endoplasmic reticulum (ER) organization in the cytoplasm generated and maintained?
(2) What sets the intrinsic length scale of self-organized cytoplasmic patterns?
(3) How do self-organized MT and ER structures carry out cell cycle functions?
How do cytoplasmic components assemble into biologically functional spatial architecture? To answer this question, we need to observe de novo assembly of spatially organized cytoplasm. This is not possible to do in living cells as they inherit their spatial organization from mother cells instead of building it from scratch. We have discovered that completely scrambled frog (Xenopus laevis) egg extracts, which contain all essential components of the cytoplasm at their native concentration, are able to carry out de novo assembly of spatially organized, biologically functional cytoplasm in vitro (Read more in Cheng and Ferrell, 2019). This unique property makes egg extracts an ideal model system for understanding the mechanisms of de novo cytoplasmic assembly.
Using the Xenopus egg extract system, we are exploring the functional impacts of cytoplasmic organization. We discovered that diffusion of protein-sized molecules (but not molecules much smaller or larger in size) becomes much faster in spatially organized cytoplasm compared to scrambled cytoplasm. Thus, self-organization could in principle enable a broad range of diffusion-limited biochemical reactions that would otherwise be too slow to be physiological (Read more in Huang, Cheng and Ferrell, 2022). We seek to further uncover the fundamental design principles of living cellular systems by dissecting the mechanisms of self-organization in frog egg cytoplasm. These principles will be instructive to the engineering of synthetic living systems, and may shed light on the origin of cellular life. My lab is currently investigating:
(1) How is the microtubule (MT) and endoplasmic reticulum (ER) organization in the cytoplasm generated and maintained?
(2) What sets the intrinsic length scale of self-organized cytoplasmic patterns?
(3) How do self-organized MT and ER structures carry out cell cycle functions?
We use microscopy, image processing and mathematical modeling to address the research questions.
Previous Work
Self-organization of the Cytoplasm
If cells are mechanically disrupted and their internal content completely scrambled, will they come back to life? Surprisingly, they sort of do. When frog eggs are crushed and their cytoplasm extracted undiluted, the homogenized cytoplasm regenerates cell-like compartments spontaneously (see the first movie below). These cell-like compartments can divide like cells (see the second movie below).
(Read more in Cheng and Ferrell, 2019).
If cells are mechanically disrupted and their internal content completely scrambled, will they come back to life? Surprisingly, they sort of do. When frog eggs are crushed and their cytoplasm extracted undiluted, the homogenized cytoplasm regenerates cell-like compartments spontaneously (see the first movie below). These cell-like compartments can divide like cells (see the second movie below).
(Read more in Cheng and Ferrell, 2019).
Cell Death Signaling
In a dying cell, apoptotic activity typically initiates from a single locus or a few isolated loci, and spreads across the entire cell. My postdoc work investigated the spatial propagation mechanism of apoptotic signaling in the cytoplasm. Using quantitative live imaging of cell-free Xenopus laevis egg extracts, intact eggs, and oocytes, we demonstrated that apoptotic signal is spread by constant-speed chemical waves in the cytoplasm. Our findings suggest that these waves, known as trigger waves, may be a general mechanism for biological communication over millimeter to centimeter distance scales. (Read more in Cheng and Ferrell, 2018).
In a dying cell, apoptotic activity typically initiates from a single locus or a few isolated loci, and spreads across the entire cell. My postdoc work investigated the spatial propagation mechanism of apoptotic signaling in the cytoplasm. Using quantitative live imaging of cell-free Xenopus laevis egg extracts, intact eggs, and oocytes, we demonstrated that apoptotic signal is spread by constant-speed chemical waves in the cytoplasm. Our findings suggest that these waves, known as trigger waves, may be a general mechanism for biological communication over millimeter to centimeter distance scales. (Read more in Cheng and Ferrell, 2018).
Systems Biology of Embryonic Development
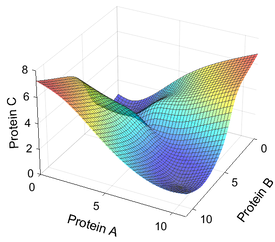
During embryogenesis, a single zygote gives rise to a multicellular embryo with distinct gene expression domains that delineate a future body plan. How is this patterning process organized? How robust is this process against perturbations? With the help of gene regulatory networks (GRNs), researchers are beginning to uncover emergent properties of developmental systems. However, the complexity of GRNs makes modeling of their dynamics a challenge. A suitable modeling framework should not only contain enough detail to recapitulate the expression dynamics, but also be computationally tractable. Combining experiments with the complementary modeling framework, we can gain a systems-level understanding of the regulatory logic not readily visible by focusing on individual genes or pathways.
(Read more in my PhD thesis).
Mathematical Modeling
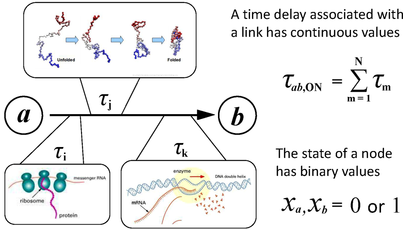
We have developed an autonomous Boolean network (ABN) modeling framework suitable for developmental regulatory systems. We show that important timing information associated with the regulatory interactions can be faithfully represented in autonomous Boolean models in which binary variables representing expression levels were updated in continuous time, and that such models can provide direct insight into features that are difficult to extract from ordinary differential equation (ODE) models. As an application, we model the experimentally well-studied network controlling fly body segmentation. The Boolean model successfully generates the patterns formed in normal and genetically perturbed fly embryos, permits the derivation of constraints on the time delay parameters, clarifies the logic associated with different ODE parameter sets, and provides a platform for studying connectivity and robustness in parameter space. By elucidating the role of regulatory time delays in pattern formation, the results suggest new types of experimental measurements in early embryonic development. (Read more in Cheng et al., 2013). We then use this framework to model the much more complicated sea urchin endomesoderm specification system. The ABN model of the sea urchin embryo captures expression patterns for several important genes during early stages of embryogenesis.
Experiments
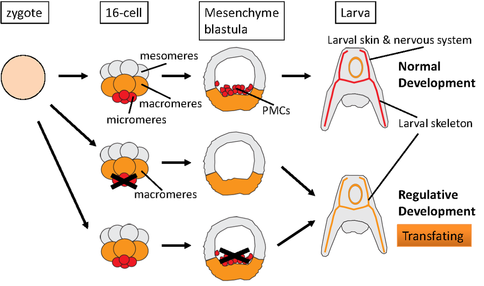
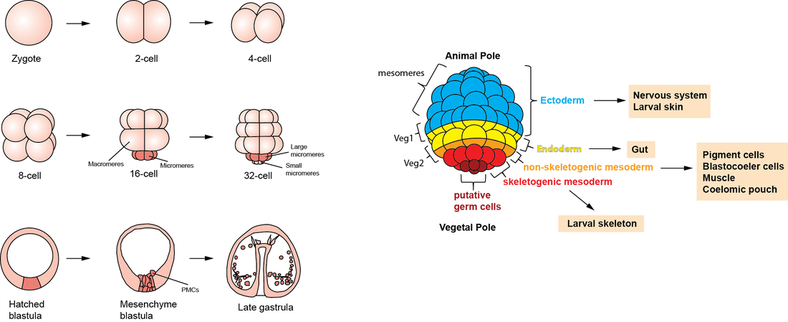
The sea urchin embryo has the remarkable ability to replace surgically removed tissues by reprogramming the presumptive fate of remaining tissues, a process known as transfating, which in turn is a form of regulative development. Understanding regulative development is essential to a complete understanding of the whole developmental regulatory program emerged from evolution. We show that regulative development requires cellular competence, and that competence is lost early on but can be regained after further differentiation. We examined micromere depletion-induced non-skeletogenic mesoderm (NSM) reprogramming and found that the skeletogenic program in NSMs undergoing reprogramming occurred within a much shorter time frame compared to normal skeletogenesis. These cells also transiently co-expressed both NSM and skeletogenic marker genes. The mesomeres normally only become ectoderm. However, isolated mesomeres are able to give rise to a complete embryo when induced by micromeres up until the 32-cell stage. Shortly after that stage, mesomeres lose the ability to be reprogrammed by transplanted micromeres. Surprisingly, in our experiments, when the mesomere descendants were isolated at late mesenchyme blastula stage, they appeared to regain the ability to give rise to a complete embryo, producing not only ectoderm, which they would normally become, but also the entirety of endoderm and mesoderm. This remarkable reprogramming potential did not require any transplanted micromeres. Further investigation revealed that this reprogramming ability depends crucially on some key transcription factors in the endomesoderm gene regulatory network. (Read more in Cheng et al., 2014).